|
Click the green Add Primer button, the Add Annotation window will open, you will see the calculated characteristics of the primer, including the Tm range based on the degeneracy, if you are happy with the primer characteristics, give the primer a Name:, go OK and the primer will be added to the consensus as an annotation.ĥ. If you are using an earlier version of Geneious, select the region, view the Statistics side pane to see the estimated Rough Tm, then go Primers -> Characteristics for Selection to add a new primer.Ĥ. If you are using Geneious Prime 2019 you will see a Floating Tooltip which details the calculated Tm for the selected region, and an Add Primer button (boxed red). On the Consensus line in the alignment, select the region you want to define as the primer, drag/select left to right for a forward primer, drag/select right to left for a reverse primer. Look for regions with minimal differences, and if possible a perfectly conserved "clamp" (as long as possible) at the 3' end of the primer.ģ. Visually identify suitable regions for a degenerate primer location using the Identity graph as a guide, zoom in and identify ideal regions for degenerate primer locations. Make sure Highlighting is on to easily see differences.Ģ. Select the alignment, make sure the consensus is on and the Threshold is set appropriately for the level of degeneracy you require. You can also do degenerate primer design manually using the following procedure.ġ. However, be aware that it will allow degeneracy very near to the 3' ends of the primers which may significantly reduce their ability to prime efficiently, even if the overall degeneracy is low. The Design New Primers tool may be suitable for your needs. With this setting, at sites where 9 out of 10 sequences are identical the most common base will be used, rather than adding a degenerate base for the single variant nucleotide. If you are using a larger alignment, and want your degeneracy to cover bases found in 90% of the sequences, set the Consensus threshold to 90% rather than 100%. 
You should now be connected to the Geneious Server.Geneious Prime can design degenerate primers based on an alignment using the Design New primers tool under the Primers menu.įor example, if you want a degenerate primer pair with primer Tm's of around 54☌, a minimum PCR product length of 150 bp (alignment is 684 bp long), and a maximum allowed degeneracy of 32 then you would use settings similar to those shown below.Ĭheck the Option to Allow Degeneracy and set an appropriate value (see the online manual, section 13.1.6 above for an explanation of degeneracy).Įnsure the option Design primers on: Consensus is set.Ĭlick the Consensus Options button and set the threshold to 100% - identical if you want the primer degeneracy to incorporate every variant base. Enter your Ceres password immediately followed by your Ceres GA code in the “Password” box as an example if your password was and GA app is showing “123456” you would use Click “OK”.Enter your Ceres username in the “User Name” box, usually firstname.lastname.You can leave the port empty, it should fill in 443 on its own. On the “Login to Geneious Server” popup screen:.You will see a message on the right “you are not currently logged in to Geneious Server”.Geneious will need to be restarted after installing new plugins.If in doubt update everything to current. It should also be noted that the Geneious Server plugin version must match the version of the Geneious Server itsself.Download the plugin then click on the file and it should install itself.If you dont see “Geneious Server” in the list of “Sources” you may need to download the “Geneious Server Plugin” from the Geneious Downloads Page.Click on “Geneious Server” in the list of Sources on the left.enter in the “License Server” box, andĪfter Geneious is started (see picture below):.On the “Enter Your License Details” screen, Geneious will complain about not having a license. 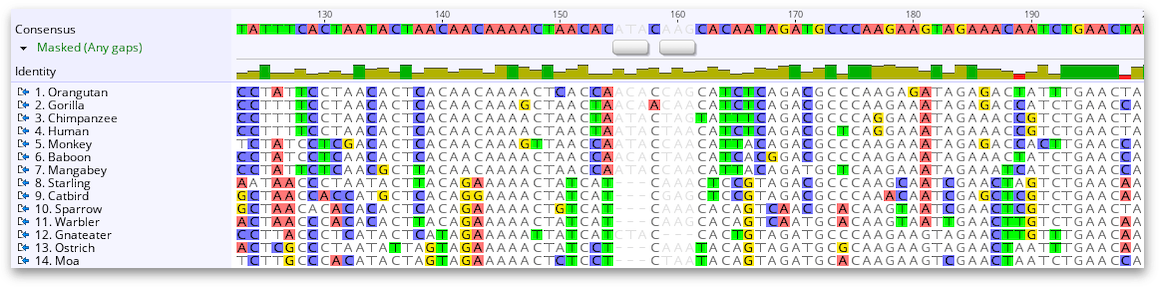 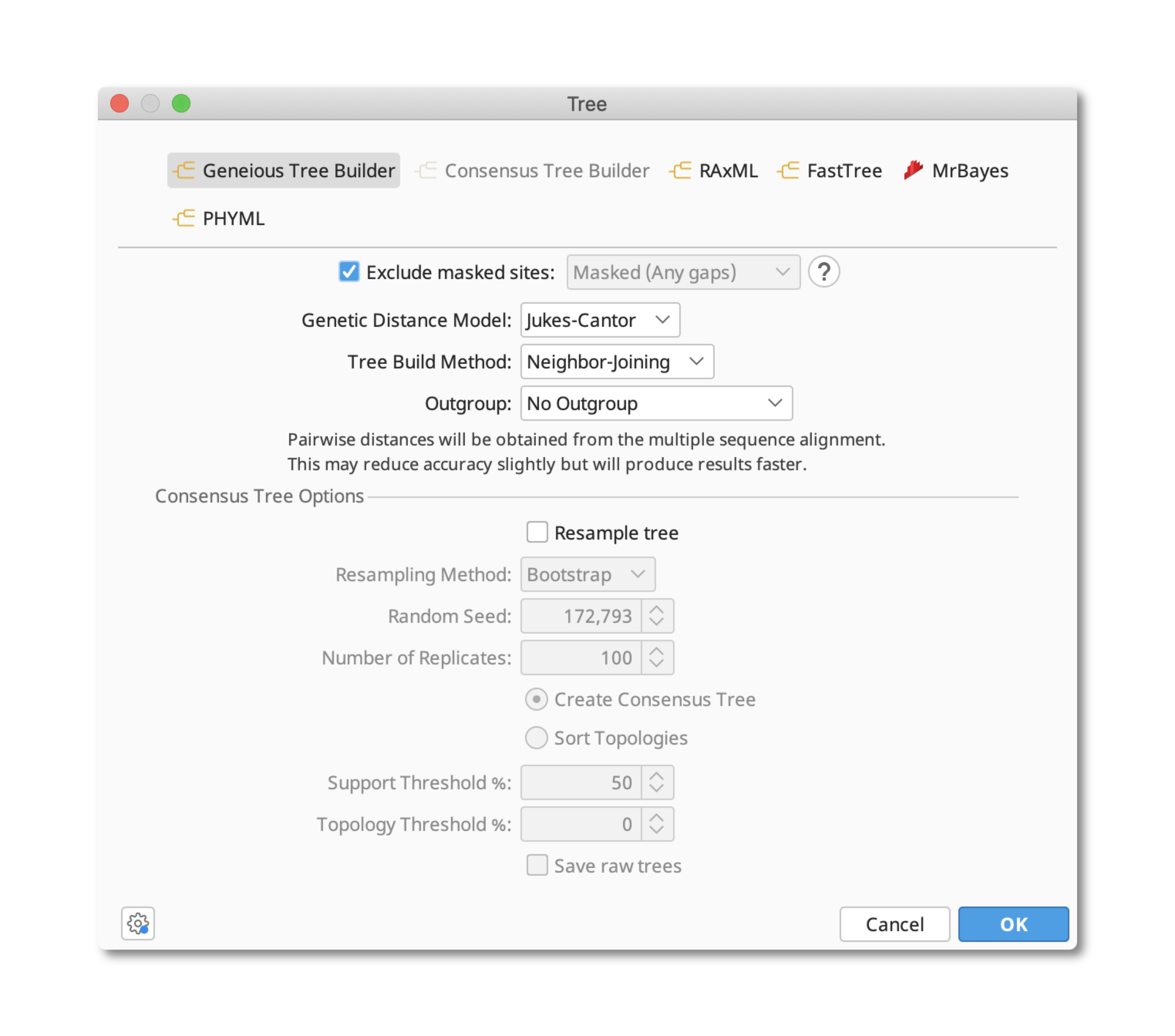
If you do encounter license server issues let us know at and start Geneious.Geneious and its license server should work through both the regular USDA VPN and the SCINet ocvpn vpn servers.The floating license server will only work at USDA sites or via VPN due to firewall restrictions.This server is also providing 20 floating licenses for Ceres users use.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
